How QTL-seq Combines Bulk Segregant Analysis and Next-Generation Sequencing
Along with classical QTL mapping in biparental populations—where breeders phenotype and genotype every individual—there has always been a powerful shortcut: Bulk Segregant Analysis (BSA).
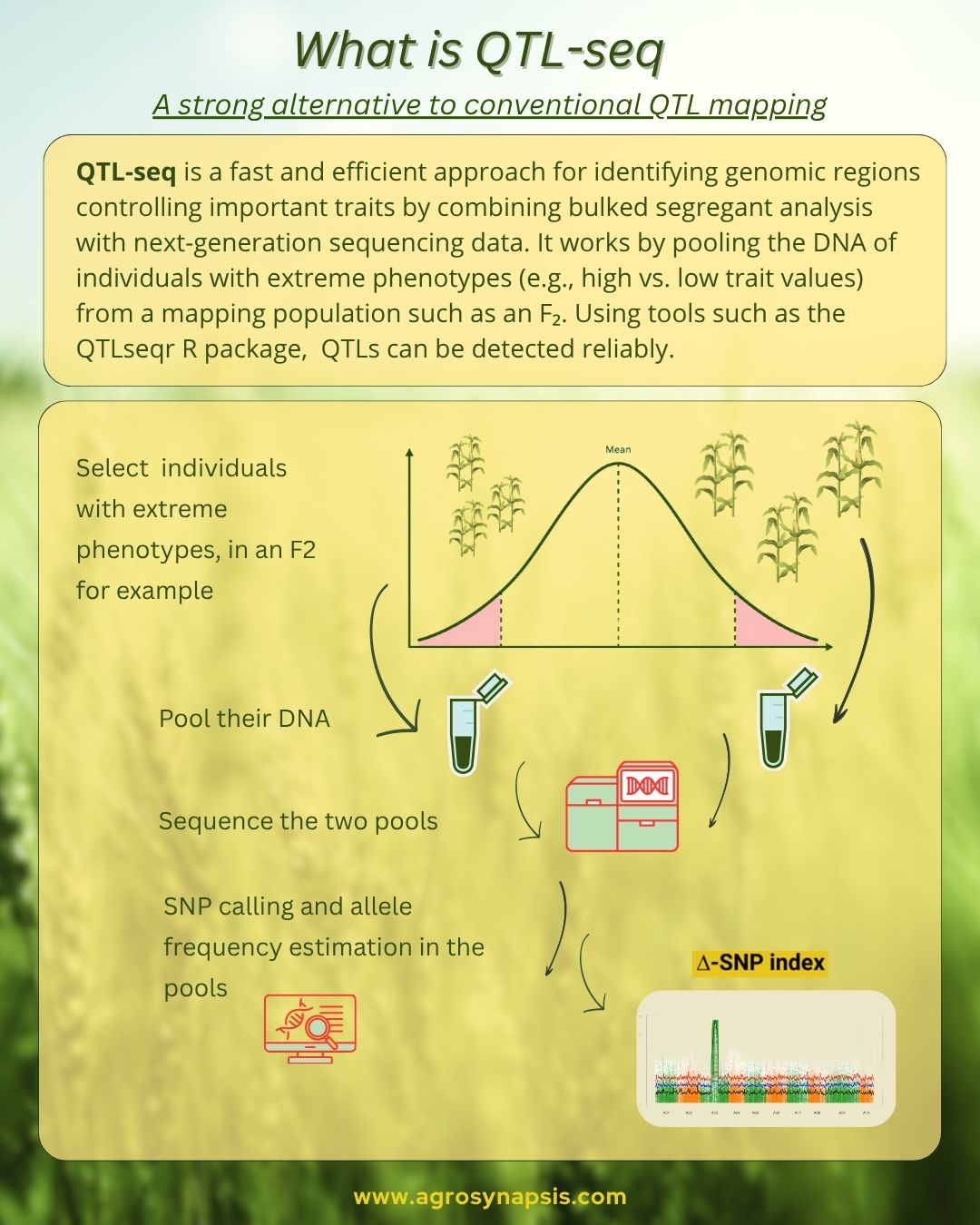
Instead of analyzing all individuals separately, BSA pools DNA from individuals showing extreme phenotypes, for example highly resistant versus highly susceptible plants, and compares allele frequencies between these contrasting groups.
Today, thanks to the dramatic reduction in sequencing costs, researchers no longer genotype these pools with only a few markers; they sequence them. This integration of BSA with next-generation sequencing (NGS) gave rise to QTL-seq.
Why QTL-seq Matters in Modern Plant Breeding
The combination of Bulk Segregant Analysis and next-generation sequencing has transformed trait mapping because QTL-seq offers:
- high mapping resolution and accuracy
- much faster analysis compared to classical QTL mapping
- significantly lower costs
- high-throughput and scalable workflows
As a result, researchers can rapidly identify genomic regions controlling important agronomic traits with precision often comparable to traditional linkage mapping approaches.
For breeding companies, this combination of speed, precision, and cost-efficiency creates a major competitive advantage, especially when studying major-effect QTLs such as disease resistance genes and when robust phenotyping protocols clearly discriminate resistant and susceptible individuals.
How QTL-seq Works in Practice
The QTL-seq workflow is elegant, efficient, and highly practical for breeding programs.
Step 1: Create a Segregating Population
Researchers first develop a segregating population, such as an F2 population generated from parents with contrasting phenotypes.
Step 2: Select Individuals with Extreme Phenotypes
Individuals showing the most contrasting phenotypes are selected, for example:
- most resistant plants
- most susceptible plants
Step 3: Construct DNA Bulks
DNA from the selected individuals is pooled into contrasting bulks:
- resistant bulk
- susceptible bulk
Step 4: Sequence the Bulks and Analyze SNP Frequencies
Researchers sequence both bulks, align the reads to a reference genome, and identify SNP markers. Next, they compare allele frequencies between the bulks by calculating the SNP-index and ΔSNP-index, often using tools such as the R package QTLseqr.
Step 5: Identify QTL Regions
Finally, genomic regions showing strong allele frequency differences between the bulks are identified as candidate QTL regions linked to the trait.
The biological principle behind QTL-seq is straightforward: marker alleles linked to the causal locus become enriched in one bulk, while neutral alleles remain randomly distributed.
Applications of QTL-seq in Crop Breeding
QTL-seq is no longer a theoretical approach. Researchers already apply it successfully across many crops and breeding objectives, including:
- disease resistance in groundnut
- fruit quality traits in tomato
- flowering time and plant height in cereals
- abiotic stress tolerance including heat, cold, and salinity
Multiple studies have successfully identified major QTLs and candidate genes using this strategy.
Practical References and Further Reading
For readers interested in exploring the methodology and practical applications of QTL-seq in greater depth, the following review provides an excellent overview:
Review on QTL-seq and Bulk Segregant Analysis
Learn More About QTL Mapping and Molecular Breeding
AgroSynapsis regularly organizes workshops and training activities focused on QTL mapping, molecular breeding, linkage mapping, and genomic data analysis.
👉 If you would like to receive workshop updates, early access, and special discounts, fill out our short training interest form: